X-rays size up protein structure at the “heart” of COVID-19 virus
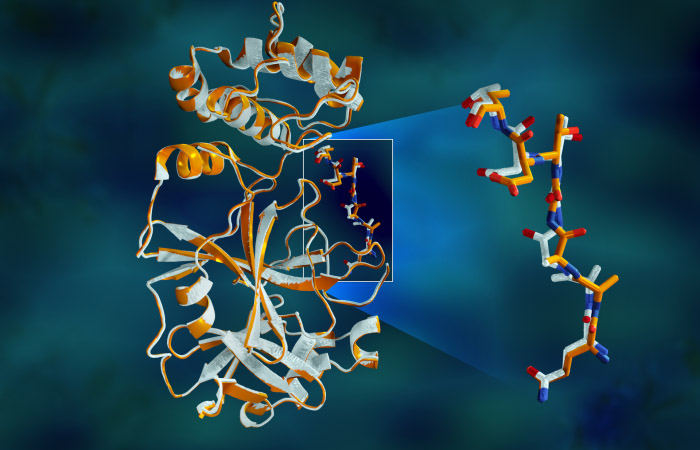
Overlapping X-ray data of the SARS-CoV-2 main protease shows structural differences between the protein at room temperature (orange) and the cryogenically frozen structure (white). Graphic: Jill Hemman/ORNL, U.S. Dept. of Energy
A team of researchers at the Department of Energy’s Oak Ridge and Argonne national laboratories has performed the first room-temperature X-ray measurements on the SARS-CoV-2 main protease, the enzyme that enables the virus to reproduce.
The X-ray measurements mark an important first step in the researchers’ ultimate goal of building a comprehensive 3D model of the enzymatic protein.